Iceberg Storage¶
Overview¶
Apache Iceberg provides the table format for feature tables, with MinIO as the S3-compatible storage backend. Together they deliver ACID transactions, time-travel queries, schema evolution, and snapshot management.
System Design | Spark Processing
Table Organization¶
Catalog Structure¶
All feature tables are organized under a single Iceberg catalog using a Hadoop-compatible backend:
quantum_catalog -- Iceberg catalog
└── quantum_features -- Database
├── molecules -- Table
├── ansatz_info -- Table
├── performance_metrics -- Table
├── vqe_results -- Table
├── initial_parameters -- Table
├── optimal_parameters -- Table
├── vqe_iterations -- Table
├── iteration_parameters -- Table
├── hamiltonian_terms -- Table
└── processing_metadata -- Table
Catalog Configuration¶
The Iceberg catalog is configured during Spark session creation:
.config('spark.sql.catalog.quantum_catalog',
'org.apache.iceberg.spark.SparkCatalog')
.config('spark.sql.catalog.quantum_catalog.type', 'hadoop')
.config('spark.sql.catalog.quantum_catalog.warehouse',
's3a://local-features/warehouse/')
| Configuration | Value | Description |
|---|---|---|
| Catalog name | quantum_catalog |
Identifier used in SQL queries |
| Catalog type | hadoop |
Hadoop-compatible catalog (metadata stored in the warehouse path) |
| Warehouse | s3a://local-features/warehouse/ |
Root location for all table data and metadata in MinIO |
Physical Storage Layout¶
On MinIO, the warehouse directory contains both metadata and data files:
s3a://local-features/warehouse/
└── quantum_features/
├── vqe_results/
│ ├── metadata/
│ │ ├── v1.metadata.json
│ │ ├── v2.metadata.json
│ │ ├── snap-1234567890.avro
│ │ └── snap-1234567891.avro
│ └── data/
│ ├── processing_date=2025-01-10/
│ │ ├── part-00000.parquet
│ │ └── part-00001.parquet
│ └── processing_date=2025-01-11/
│ └── part-00000.parquet
├── vqe_iterations/
│ ├── metadata/
│ └── data/
└── molecules/
├── metadata/
└── data/
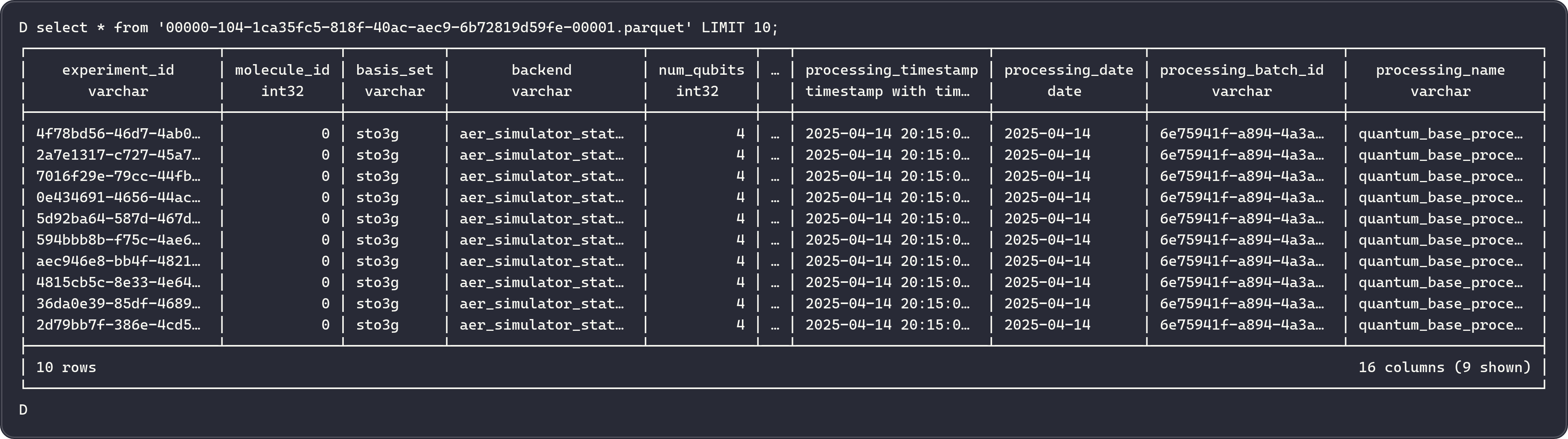
Metadata Files¶
| File Type | Description |
|---|---|
v*.metadata.json |
Table schema, partition spec, current snapshot pointer, and table properties |
snap-*.avro |
Snapshot metadata containing manifest list references |
| Manifest lists | Lists of manifest files for a given snapshot |
| Manifest files | Lists of data files, their partition values, and file-level statistics |
Iceberg Table Features¶
Iceberg provides ACID transactions, time-travel queries, schema evolution, and snapshot management without requiring data rewrites. For full details on these capabilities, see the Apache Iceberg documentation.
Snapshot Tagging¶
Each Spark write creates a new snapshot, tagged with a version identifier from the processing batch ID:
snapshot_id = spark.sql(
f'SELECT snapshot_id '
f'FROM quantum_catalog.quantum_features.{table_name}.snapshots '
f'ORDER BY committed_at DESC LIMIT 1'
).collect()[0][0]
version_tag = f'v_{processing_batch_id}'
spark.sql(f"""
ALTER TABLE quantum_catalog.quantum_features.{table_name}
CREATE TAG {version_tag} AS OF VERSION {snapshot_id}
""")
Reproducible ML Training
Reference a snapshot tag from a specific processing run to reproduce exact training datasets.
MinIO Integration¶
MinIO provides S3-compatible storage for raw Avro data and processed Parquet feature tables.
Bucket Structure¶
The system uses two MinIO buckets:
| Bucket | Purpose | Writer |
|---|---|---|
local-vqe-results |
Raw Avro files from Kafka Connect | Kafka Connect S3 Sink |
local-features |
Processed Parquet files and Iceberg metadata | Apache Spark |
Kafka Connect S3 Sink¶
Kafka Connect writes raw Avro files to MinIO via the S3 Sink Connector:
{
"connector.class": "io.confluent.connect.s3.S3SinkConnector",
"tasks.max": "1",
"topics.regex": "vqe_decorated_result_.*",
"topics.dir": "experiments",
"s3.bucket.name": "local-vqe-results",
"store.url": "http://minio:9000",
"s3.region": "us-east-1",
"s3.path.style.access": "true",
"flush.size": "1",
"schema.compatibility": "NONE",
"storage.class": "io.confluent.connect.s3.storage.S3Storage",
"format.class": "io.confluent.connect.s3.format.avro.AvroFormat"
}
Full configuration: docker/connectors/minio-sink-config.json
Raw Data Directory Layout¶
s3://local-vqe-results/
└── experiments/
├── vqe_decorated_result_v1/
│ ├── partition=0/
│ │ ├── vqe_decorated_result_v1+0+0000000000.avro
│ │ ├── vqe_decorated_result_v1+0+0000000001.avro
│ │ └── vqe_decorated_result_v1+0+0000000002.avro
│ └── partition=1/
│ └── vqe_decorated_result_v1+1+0000000000.avro
└── vqe_decorated_result_v2/
└── partition=0/
└── vqe_decorated_result_v2+0+0000000000.avro
File naming convention: {topic_name}+{partition}+{start_offset}.avro
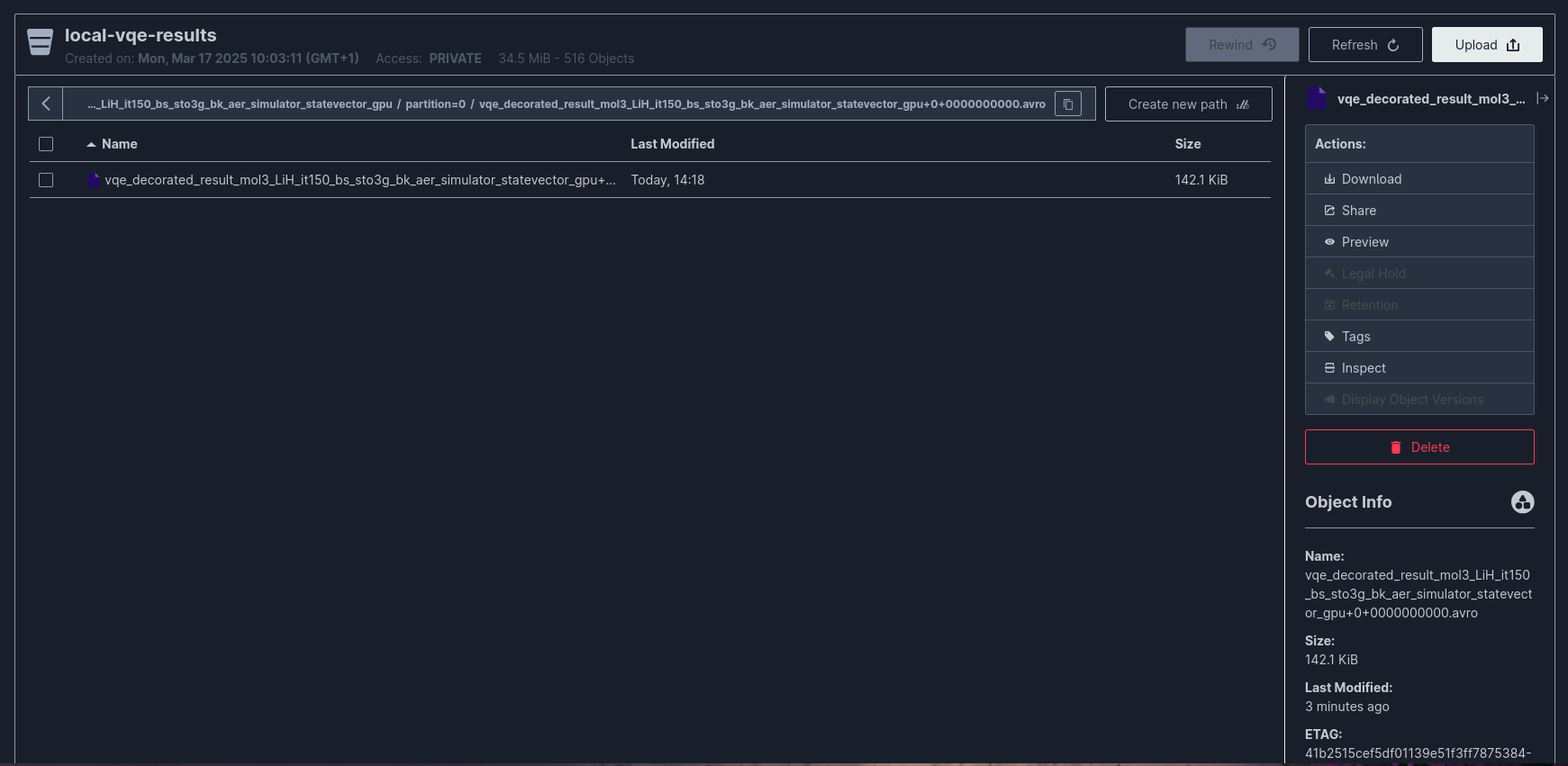
MinIO Configuration¶
minio:
image: minio/minio:${MINIO_VERSION:-latest}
command: minio server /data --console-address ":${MINIO_CONSOLE_PORT}"
ports:
- "${MINIO_API_PORT}:9000" # S3 API
- "${MINIO_CONSOLE_PORT}:9001" # Web Console
environment:
MINIO_ROOT_USER: ${MINIO_ROOT_USER}
MINIO_ROOT_PASSWORD: ${MINIO_ROOT_PASSWORD}
MINIO_REGION_NAME: ${MINIO_REGION}
volumes:
- minio-data:/data
| Port | Service | Description |
|---|---|---|
${MINIO_API_PORT} |
S3 API | Used by Kafka Connect and Spark for data access |
${MINIO_CONSOLE_PORT} |
Web Console | Browser-based management interface |
Partitioning Strategy¶
Partitioning is optimized for expected query patterns:
| Table | Partition Columns | Rationale |
|---|---|---|
molecules |
processing_date |
Time-based filtering for incremental loads |
ansatz_info |
processing_date, basis_set |
Filter by date and basis set for circuit analysis |
performance_metrics |
processing_date, basis_set |
Performance comparisons across basis sets |
vqe_results |
processing_date, basis_set, backend |
Query by date, basis set, and simulation backend |
initial_parameters |
processing_date, basis_set |
Parameter analysis by date and basis set |
optimal_parameters |
processing_date, basis_set |
Optimized parameter lookup |
vqe_iterations |
processing_date, basis_set, backend |
Iteration analysis with backend filtering |
iteration_parameters |
processing_date, basis_set |
Per-iteration parameter tracking |
hamiltonian_terms |
processing_date, basis_set, backend |
Hamiltonian structure analysis |
Partition Pruning¶
Iceberg uses partition metadata to skip irrelevant data files at query time:
-- Only scans partitions for sto3g basis set on a specific date
SELECT * FROM quantum_catalog.quantum_features.vqe_results
WHERE processing_date = DATE '2025-01-12'
AND basis_set = 'sto3g';
Iceberg reads only files in matching partitions, reducing I/O for partition-filtered queries.
Table Maintenance¶
Iceberg tables require periodic maintenance including snapshot expiration, data file compaction, and orphan file cleanup. For procedures and recommended schedules, see the Iceberg Maintenance documentation.
Processing Metadata Table¶
A metadata table tracks all processing runs:
CREATE TABLE IF NOT EXISTS quantum_catalog.quantum_features.processing_metadata (
processing_batch_id STRING,
processing_name STRING,
processing_timestamp TIMESTAMP,
processing_date DATE,
table_names ARRAY<STRING>,
table_versions ARRAY<STRING>,
record_counts ARRAY<BIGINT>,
source_data_info STRING
) USING iceberg
This provides an audit trail for tracing which data was processed and when.
Related Documentation¶
- System Design - Full architecture overview
- Spark Processing - Feature engineering pipeline that writes to Iceberg
- Kafka Streaming - How raw data arrives in MinIO via Kafka Connect
- Airflow Orchestration - Scheduling of the processing pipeline