Spark Processing¶
Overview¶
Apache Spark transforms raw VQE simulation results into structured ML feature tables. The processing is batch-oriented and incremental - only new data since the last run is processed.
System Design | Airflow Orchestration
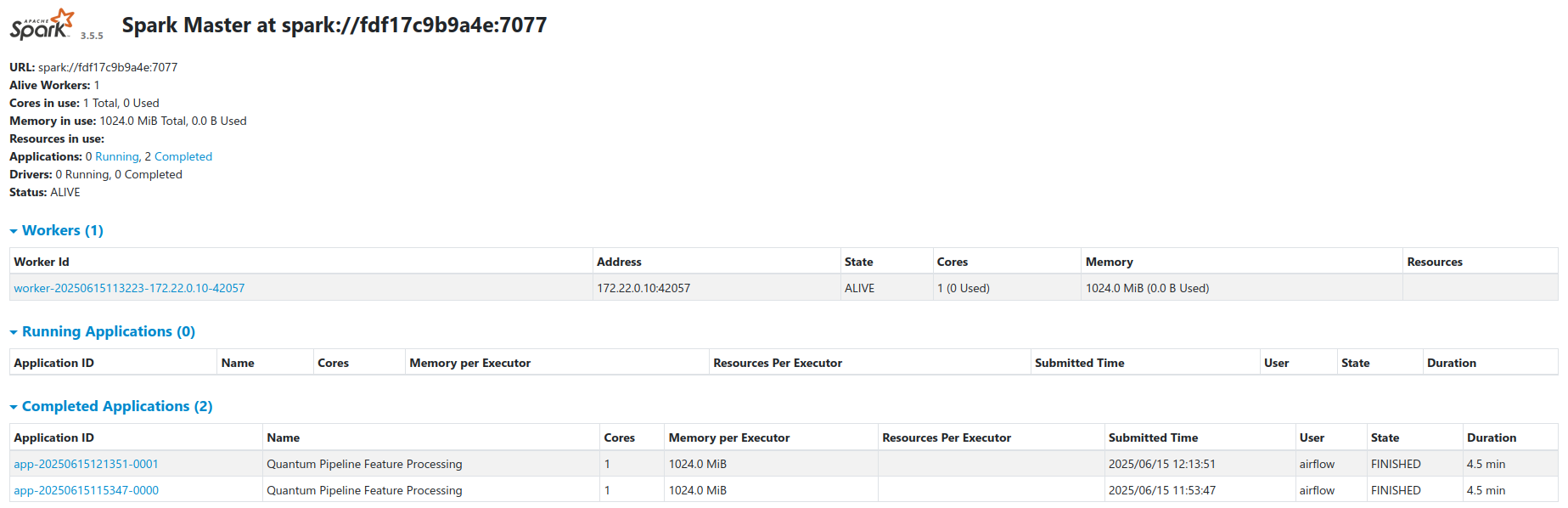
Cluster Architecture¶
The cluster runs in standalone master-worker mode via a custom Spark Docker image (docker/Dockerfile.spark). See the Spark documentation for general cluster configuration.
| Node | Role | Resources |
|---|---|---|
spark-master |
Coordinator | Default (no explicit limits) |
spark-worker |
Executor | 1 core, 1 GB RAM (docker-compose.yaml) |
The thesis compose file (docker-compose.thesis.yaml) increases the worker to 2 cores and 4 GB RAM with explicit resource limits.
- Workers register with the master at
spark://spark-master:7077 - Airflow submits jobs via the
SparkSubmitOperator - Workers access MinIO directly through the S3A filesystem connector
- Spark Web UI is available at port
8080on the master node
Feature Engineering Pipeline¶
A 6-step workflow transforms raw VQE results into nine feature tables. Each run is incremental.
Step 1: Create Spark Session¶
A SparkSession is initialized with S3A (MinIO connectivity), Iceberg catalog integration, and adaptive query execution. The configuration is defined in DEFAULT_CONFIG at the top of the processing script.
spark = (
SparkSession.builder
.appName(config.get('APP_NAME'))
.master(config.get('SPARK_MASTER'))
.config('spark.sql.extensions',
'org.apache.iceberg.spark.extensions.IcebergSparkSessionExtensions')
.config('spark.sql.catalog.quantum_catalog',
'org.apache.iceberg.spark.SparkCatalog')
.config('spark.sql.catalog.quantum_catalog.type', 'hadoop')
.config('spark.sql.catalog.quantum_catalog.warehouse',
config.get('S3_WAREHOUSE'))
.config('spark.hadoop.fs.s3a.impl',
'org.apache.hadoop.fs.s3a.S3AFileSystem')
.config('spark.hadoop.fs.s3a.endpoint', config.get('S3_ENDPOINT'))
.config('spark.hadoop.fs.s3a.path.style.access', 'true')
.config('spark.sql.adaptive.enabled', 'true')
.config('spark.sql.shuffle.partitions', '200')
.getOrCreate()
)
Default S3 paths from DEFAULT_CONFIG:
- Experiment bucket:
s3a://local-vqe-results/experiments/ - Feature warehouse:
s3a://local-features/warehouse/
Step 2: Initialize Iceberg Metadata¶
The job reads or initializes the Iceberg metadata catalog. If this is the first run, the catalog and database are created. On subsequent runs, existing metadata is loaded to determine which data has already been processed.
def list_available_topics(spark, bucket_path):
"""List available topics from the storage."""
fs = spark._jvm.org.apache.hadoop.fs.FileSystem.get(
spark._jvm.java.net.URI.create(bucket_path),
spark._jsc.hadoopConfiguration()
)
path = spark._jvm.org.apache.hadoop.fs.Path(bucket_path)
if fs.exists(path) and fs.isDirectory(path):
return [f.getPath().getName() for f in fs.listStatus(path)
if f.isDirectory()]
return []
Step 3: Filter New Data¶
New records are identified by joining against existing Iceberg table keys. A marker-column approach is used instead of left_anti to handle edge cases.
def identify_new_records(spark, new_data_df, table_name, key_columns):
"""Identify records not yet present in the target Iceberg table."""
if not spark.catalog.tableExists(
f'quantum_catalog.quantum_features.{table_name}'):
return new_data_df
existing_keys = spark.sql(
f'SELECT DISTINCT {", ".join(key_columns)} '
f'FROM quantum_catalog.quantum_features.{table_name}'
)
if existing_keys.isEmpty():
return new_data_df
# Marker-column join to find only new records
new_with_marker = (new_data_df.select(*key_columns).distinct()
.withColumn('is_new', lit(1)))
existing_with_marker = existing_keys.withColumn('exists', lit(1))
joined = new_with_marker.join(existing_with_marker,
on=key_columns, how='left')
new_keys = joined.filter(col('exists').isNull()).select(*key_columns)
return new_data_df.join(new_keys, on=key_columns, how='inner')
Step 4: Extract Features¶
Raw VQE data is transformed into nine specialized feature tables. The transform_quantum_data function flattens nested Avro fields and adds metadata columns (experiment_id, processing_timestamp, processing_date, processing_batch_id).
def transform_quantum_data(df):
"""Transform raw VQE data into specialized feature tables."""
base_df = add_metadata_columns(df, 'quantum_base_processing')
# Flatten nested Avro fields into a wide DataFrame, then
# select columns for each feature table:
return {
'molecules': df_molecule,
'ansatz_info': df_ansatz,
'performance_metrics': df_metrics,
'vqe_results': df_vqe,
'initial_parameters': df_initial_parameters,
'optimal_parameters': df_optimal_parameters,
'vqe_iterations': df_iterations,
'iteration_parameters': df_iteration_parameters,
'hamiltonian_terms': df_hamiltonian,
}
See docker/airflow/scripts/quantum_incremental_processing.py for the full implementation of each extraction.
Step 5: Write Parquet to MinIO¶
Feature data is written as Parquet through the Iceberg table API. New tables are created on first run; subsequent runs append only new records.
def process_incremental_data(spark, new_data_df, table_name, key_columns,
partition_columns=None):
"""Write new data to an Iceberg table, creating it if necessary."""
table_path = f'quantum_catalog.quantum_features.{table_name}'
if not spark.catalog.tableExists(table_path):
writer = new_data_df.write.format('iceberg') \
.option('write-format', 'parquet')
if partition_columns:
writer = writer.partitionBy(*partition_columns)
writer.mode('overwrite').saveAsTable(table_path)
else:
truly_new = identify_new_records(spark, new_data_df,
table_name, key_columns)
if truly_new.count() > 0:
truly_new.write.format('iceberg') \
.option('write-format', 'parquet') \
.mode('append') \
.saveAsTable(table_path)
Step 6: Update Iceberg Snapshots¶
After writing, a tagged Iceberg snapshot records the processing batch ID for time-travel queries. Initial writes use v_{batch_id} tags; incremental appends use v_incr_{batch_id}.
snapshot_id = spark.sql(
f'SELECT snapshot_id '
f'FROM quantum_catalog.quantum_features.{table_name}.snapshots '
f'ORDER BY committed_at DESC LIMIT 1'
).collect()[0][0]
version_tag = f'v_{processing_batch_id}'
spark.sql(f"""
ALTER TABLE quantum_catalog.quantum_features.{table_name}
CREATE TAG {version_tag} AS OF VERSION {snapshot_id}
""")
Feature Tables¶
Nine tables stored as Iceberg-managed Parquet files in MinIO under quantum_catalog.quantum_features.
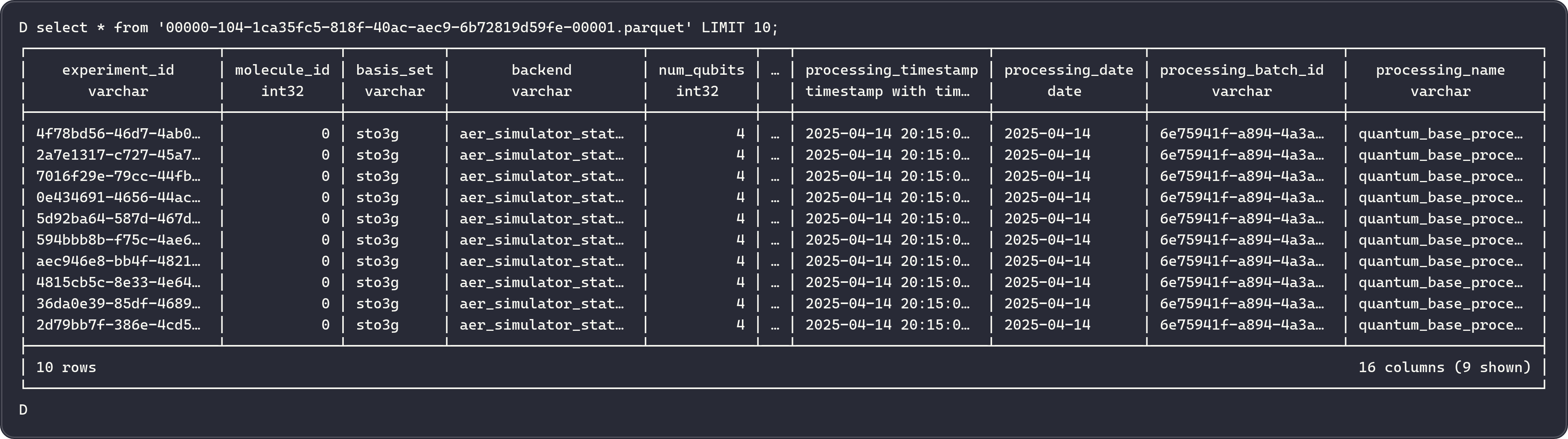
1. molecules¶
Molecular structure information for each simulated molecule.
| Column | Type | Description |
|---|---|---|
experiment_id |
string | Unique experiment identifier |
molecule_id |
int | Molecule identifier |
atom_symbols |
array<string> | Chemical symbols (e.g., ["H", "H"]) |
coordinates |
array<array<double>> | 3D atomic coordinates |
multiplicity |
int | Spin multiplicity |
charge |
int | Molecular charge |
coordinate_units |
string | Units (angstrom or bohr) |
atomic_masses |
array<double> | Atomic masses (amu) |
Partitioned by: processing_date
2. ansatz_info¶
Quantum circuit ansatz configurations used in each experiment.
| Column | Type | Description |
|---|---|---|
experiment_id |
string | Unique experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set (e.g., sto3g, cc-pvdz) |
ansatz |
string | QASM3 circuit representation |
ansatz_reps |
int | Number of ansatz repetitions |
Partitioned by: processing_date, basis_set
3. performance_metrics¶
Execution timing and performance data for each simulation.
| Column | Type | Description |
|---|---|---|
experiment_id |
string | Unique experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
hamiltonian_time |
double | Hamiltonian construction time (seconds) |
mapping_time |
double | Fermionic-to-qubit mapping time (seconds) |
vqe_time |
double | VQE optimization time (seconds) |
total_time |
double | Total simulation time (seconds) |
minimization_time |
double | Optimizer execution time (seconds) |
computed_total_time |
double | Sum of hamiltonian_time + mapping_time + vqe_time |
Partitioned by: processing_date, basis_set
4. vqe_results¶
Core VQE optimization results including the minimum energy found.
| Column | Type | Description |
|---|---|---|
experiment_id |
string | Unique experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
backend |
string | Qiskit backend (e.g., aer_simulator) |
num_qubits |
int | Number of qubits in the circuit |
optimizer |
string | Optimizer algorithm (e.g., L-BFGS-B, COBYLA) |
noise_backend |
string | Noise model used (if any) |
default_shots |
int | Number of shots per circuit execution |
ansatz_reps |
int | Number of ansatz repetitions |
minimum_energy |
double | Ground state energy estimate (Hartree) |
maxcv |
double | Maximum constraint violation |
total_iterations |
int | Number of optimizer iterations |
Partitioned by: processing_date, basis_set, backend
5. initial_parameters¶
Starting variational parameters for each experiment.
| Column | Type | Description |
|---|---|---|
parameter_id |
string | Unique parameter identifier |
experiment_id |
string | Experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
backend |
string | Qiskit backend |
num_qubits |
int | Number of qubits |
parameter_index |
int | Parameter position in the circuit |
initial_parameter_value |
double | Starting parameter value |
Partitioned by: processing_date, basis_set
6. optimal_parameters¶
Optimized variational parameters after VQE convergence.
| Column | Type | Description |
|---|---|---|
parameter_id |
string | Unique parameter identifier |
experiment_id |
string | Experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
backend |
string | Qiskit backend |
num_qubits |
int | Number of qubits |
parameter_index |
int | Parameter position in the circuit |
optimal_parameter_value |
double | Optimized parameter value |
Partitioned by: processing_date, basis_set
7. vqe_iterations¶
Per-iteration optimization data recording the energy at each step.
| Column | Type | Description |
|---|---|---|
iteration_id |
string | Unique iteration identifier |
experiment_id |
string | Experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
backend |
string | Qiskit backend |
num_qubits |
int | Number of qubits |
iteration_step |
int | Iteration number |
iteration_energy |
double | Energy at this step (Hartree) |
energy_std_dev |
double | Standard deviation of measurement |
Partitioned by: processing_date, basis_set, backend
8. iteration_parameters¶
Parameter values at each optimizer iteration, enabling convergence analysis.
| Column | Type | Description |
|---|---|---|
parameter_id |
string | Unique parameter identifier |
iteration_id |
string | Iteration identifier |
experiment_id |
string | Experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
backend |
string | Qiskit backend |
num_qubits |
int | Number of qubits |
iteration_step |
int | Iteration number |
parameter_value |
double | Parameter value at this iteration |
Partitioned by: processing_date, basis_set
9. hamiltonian_terms¶
Hamiltonian Pauli operator terms and their complex coefficients.
| Column | Type | Description |
|---|---|---|
term_id |
string | Unique term identifier |
experiment_id |
string | Experiment identifier |
molecule_id |
int | Molecule identifier |
basis_set |
string | Basis set |
backend |
string | Qiskit backend |
term_label |
string | Pauli string (e.g., "XYZI", "IIZZ") |
coeff_real |
double | Real part of the coefficient |
coeff_imag |
double | Imaginary part of the coefficient |
Partitioned by: processing_date, basis_set, backend
Incremental Processing¶
Spark uses Iceberg metadata to identify and process only new records on each run.
How It Works¶
- List current files - Spark enumerates all Avro files in the MinIO experiments bucket.
- Check Iceberg snapshots - If an Iceberg snapshot exists from a previous run, the system retrieves the set of already-processed file paths.
- Compute delta - New files are identified as the difference between current files and previously processed files.
- Process delta only - Only the new files are read, transformed, and appended to the feature tables.
- Update snapshot - A new Iceberg snapshot is created, recording the current set of processed files.
Related Documentation¶
- System Design - Full architecture overview
- Airflow Orchestration - Spark job scheduling
- Iceberg Storage - Table format and snapshots
- Kafka Streaming - Raw data ingestion