System Design Documentation¶
Overview¶
This chapter presents the architecture of the system designed for simulating and processing Variational Quantum Eigensolver (VQE) algorithm results.
It consists of the following key components:
- Simulation module
- Apache Kafka message broker
- MinIO storage system
- Apache Airflow orchestrator
- Apache Spark processing engine (master-worker configuration with one worker)
Each component communicates with the others through a Docker container network, ensuring isolation, consistency, and the ability to scale independently. The architecture is based on the microservices pattern.
Architecture Principles¶
The system follows several key architectural principles:
- Separation of Concerns: Each component has a well-defined responsibility
- Loose Coupling: Components communicate through well-defined interfaces
- Scalability: Horizontal scaling capabilities for compute-intensive tasks
- Data Versioning: Immutable data with schema evolution support
- Incremental Processing: Only process new data to optimize resources
- Observability: Comprehensive monitoring and metrics collection
1. Quantum Simulation Module¶
Overview¶
The main simulation module prepares data for simulation and executes VQE simulations using the Qiskit Aer simulator. The system was designed with the capability to integrate with real quantum computers via IBM Quantum, but due to access costs, only the simulator is extensively tested and used in this project.
The VQE algorithm is monitored during execution - data is collected from each optimizer iteration, execution times for individual stages, and information about the molecule and the basis set used.
Each simulation module container also exports system metrics (CPU and RAM usage) to Prometheus PushGateway, enabling resource monitoring in a containerized environment.
VQE Execution Pipeline¶
sequenceDiagram
participant User
participant VQERunner
participant QiskitAer
participant Monitor
participant Prometheus
participant Kafka
User->>VQERunner: Start VQE Simulation
VQERunner->>VQERunner: Load Molecule
VQERunner->>VQERunner: Build Hamiltonian
Note over VQERunner: Track hamiltonian_time
VQERunner->>VQERunner: Map to Qubits
Note over VQERunner: Track mapping_time
VQERunner->>Monitor: Start Performance Monitoring
Monitor->>Prometheus: Export System Metrics
VQERunner->>QiskitAer: Execute VQE
loop Each Optimizer Iteration
QiskitAer->>QiskitAer: Evaluate Cost Function
QiskitAer->>Monitor: Log Iteration Data
Monitor->>Monitor: Store Parameters & Energy
end
Note over QiskitAer: Track vqe_time
QiskitAer->>VQERunner: Return Optimization Result
VQERunner->>VQERunner: Aggregate Results
Note over VQERunner: Calculate total_time
VQERunner->>Kafka: Publish VQEDecoratedResult
Monitor->>Prometheus: Export Final Metrics
VQERunner->>User: Simulation CompleteData Collection During Execution¶
All collected data are aggregated into a single result object, which size depends on the problem complexity and the number of algorithm iterations.
The monitoring system collects:
- Iteration Data: Parameters, energy values, and standard deviation for each optimizer step
- Timing Information:
hamiltonian_time: Time to construct the molecular Hamiltonianmapping_time: Time to map fermionic operators to qubit operatorsvqe_time: VQE optimization execution timetotal_time: Total simulation time
- Molecule Information: Atomic symbols, coordinates, charge, multiplicity, basis set
- System Metrics: CPU usage, memory consumption, container resource utilization
VQEDecoratedResult Interface¶
The implementation uses Apache Avro as the data serialization format.
VQEDecoratedResultInterface class is responsible for serialization, deserialization, and managing
the data schema in the schema registry.
This separation of responsibilities ensures data format consistency between different system components. Long term planned is migration to a more readable and less cluttered Pydantic structure.
class VQEDecoratedResultInterface(AvroInterfaceBase[VQEDecoratedResult]):
def __init__(self, registry):
super().__init__(registry)
self.result_interface = VQEResultInterface(self.registry)
self.molecule_interface = MoleculeInfoInterface(self.registry)
self.schema_name = 'vqe_decorated_result'
@property
def schema(self) -> dict[str, Any]:
try:
return self.registry.get_schema(self.schema_name)
except FileNotFoundError:
schema = {
'type': 'record',
'name': 'VQEDecoratedResult',
'fields': [
{'name': 'vqe_result', 'type': self.result_interface.schema},
{'name': 'molecule', 'type': self.molecule_interface.schema},
{'name': 'basis_set', 'type': 'string'},
{'name': 'hamiltonian_time', 'type': 'double'},
{'name': 'mapping_time', 'type': 'double'},
{'name': 'vqe_time', 'type': 'double'},
{'name': 'total_time', 'type': 'double'},
{'name': 'molecule_id', 'type': 'int'},
],
}
dict_schema = deepcopy(schema)
self.registry.save_schema(self.schema_name, schema)
return dict_schema
def serialize(self, obj: VQEDecoratedResult) -> dict[str, Any]:
return {
'vqe_result': self.result_interface.serialize(obj.vqe_result),
'molecule': self.molecule_interface.serialize(obj.molecule),
'basis_set': obj.basis_set,
'hamiltonian_time': float(obj.hamiltonian_time),
'mapping_time': float(obj.mapping_time),
'vqe_time': float(obj.vqe_time),
'total_time': float(obj.total_time),
'molecule_id': obj.molecule_id,
}
def deserialize(self, data: dict[str, Any]) -> VQEDecoratedResult:
return VQEDecoratedResult(
vqe_result=self.result_interface.deserialize(data['vqe_result']), # Simulation results - including each iteration
molecule=self.molecule_interface.deserialize(data['molecule']), # Information about simulated molecule
basis_set=data['basis_set'], # Basis set - determines simulation accuracy
hamiltonian_time=float64(data['hamiltonian_time']), # Time to build Hamiltonian operator
mapping_time=float64(data['mapping_time']), # Mapping time
vqe_time=float64(data['vqe_time']), # VQE algorithm simulation time
total_time=float64(data['total_time']), # Total simulation time
molecule_id=int(data['molecule_id']), # Molecule identifier
)
Schema Structure Breakdown¶
Nested Schema Architecture
The VQEDecoratedResult schema uses a compositional design pattern, where
complex types are built from simpler nested schemas.
Top-Level Schema: VQEDecoratedResult¶
| Field | Type | Description |
|---|---|---|
vqe_result |
VQEResult (nested) | Complete VQE optimization results |
molecule |
MoleculeInfo (nested) | Molecular structure information |
basis_set |
string | Basis set used (e.g., sto3g, cc-pvdz) |
hamiltonian_time |
double | Hamiltonian construction time (seconds) |
mapping_time |
double | Fermionic-to-qubit mapping time (seconds) |
vqe_time |
double | VQE optimization execution time (seconds) |
total_time |
double | Total end-to-end execution time (seconds) |
molecule_id |
int | Unique molecule identifier |
VQEResult Schema¶
| Field | Type | Description |
|---|---|---|
initial_data |
VQEInitialData (nested) | Initial configuration and parameters |
iteration_list |
array<VQEProcess> | Complete iteration history |
minimum |
double | Minimum energy found (Hartree) |
optimal_parameters |
array<double> | Optimal circuit parameters |
maxcv |
double (nullable) | Maximum constraint violation |
minimization_time |
double | Optimizer execution time (seconds) |
VQEInitialData Schema¶
| Field | Type | Description |
|---|---|---|
backend |
string | Qiskit backend name (e.g., "aer_simulator") |
num_qubits |
int | Number of qubits in circuit |
hamiltonian |
array<HamiltonianTerm> | Pauli operator terms with coefficients |
num_parameters |
int | Number of variational parameters |
initial_parameters |
array<double> | Starting parameter values |
optimizer |
string | Optimizer name (e.g., "L-BFGS-B") |
ansatz |
string | QASM3 representation of circuit |
ansatz_reps |
int | Number of ansatz repetitions |
noise_backend |
string | Noise model configuration |
default_shots |
int | Number of measurement shots |
VQEProcess Schema (Iteration Data)¶
| Field | Type | Description |
|---|---|---|
iteration |
int | Iteration number |
parameters |
array<double> | Parameter values at this iteration |
result |
double | Energy value (Hartree) |
std |
double | Standard deviation of measurement |
MoleculeInfo Schema¶
| Field | Type | Description |
|---|---|---|
symbols |
array<string> | Atomic symbols (e.g., ["H", "H"]) |
coords |
array<array<double>> | 3D coordinates (Angstrom or Bohr) |
multiplicity |
int | Spin multiplicity |
charge |
int | Total molecular charge |
units |
string | Coordinate units ("angstrom" or "bohr") |
masses |
array<double> (nullable) | Atomic masses (amu) |
HamiltonianTerm Schema¶
| Field | Type | Description |
|---|---|---|
label |
string | Pauli string (e.g., "XYZI", "IIZZ") |
coefficients |
ComplexNumber | Complex coefficient |
ComplexNumber Schema¶
| Field | Type | Description |
|---|---|---|
real |
double | Real component |
imaginary |
double | Imaginary component |
Serialization Details¶
Type Conversion Strategy
The Avro interface handles automatic conversion between Python/NumPy types and Avro-compatible primitives:
numpy.float64->doublenumpy.int64->long/intnumpy.ndarray->array<T>- Complex numbers ->
{real: double, imaginary: double}
Binary Serialization with Schema Registry¶
After serialization, data is transmitted to Apache Kafka. The serialization process includes:
- Schema Registration: Schema is registered with Confluent Schema Registry
- Magic Byte Prefix: Binary data starts with
0x00(Confluent wire format) - Schema ID: 4-byte big-endian schema identifier
- Avro Binary Data: Compact binary-encoded payload
def to_avro_bytes(self, obj: T, schema_name: str = 'vqe_decorated_result') -> bytes:
"""Convert object to Avro binary format."""
schema = self.schema
parsed_schema = avro.schema.parse(json.dumps(schema))
writer = DatumWriter(parsed_schema)
bytes_writer = io.BytesIO()
# Write Confluent Schema Registry header
bytes_writer.write(bytes([0])) # Magic byte
bytes_writer.write(self.registry.id_cache[schema_name].to_bytes(4, byteorder='big'))
encoder = BinaryEncoder(bytes_writer)
writer.write(self.serialize(obj), encoder)
return bytes_writer.getvalue()
Container Metrics Export¶
Each simulation container exports metrics to Prometheus PushGateway:
graph LR
QC1[Quantum Container 1] -->|HTTP POST| PG[Prometheus PushGateway]
QC2[Quantum Container 2] -->|HTTP POST| PG
QC3[Quantum Container 3] -->|HTTP POST| PG
PG -->|Scrape| PROM[Prometheus]
PROM -->|Query| GRAF[Grafana Dashboards]
style QC1 fill:#c5cae9,color:#1a237e
style QC2 fill:#c5cae9,color:#1a237e
style QC3 fill:#c5cae9,color:#1a237e
style PG fill:#e8f5e9,color:#1b5e20
style PROM fill:#e8f5e9,color:#1b5e20
style GRAF fill:#e8f5e9,color:#1b5e20Exported Metrics:
quantum_system_cpu_percent: CPU utilization percentagequantum_system_memory_percent: Memory utilization percentagequantum_vqe_total_time: Total VQE execution timequantum_vqe_minimum_energy: Minimum energy value foundquantum_vqe_iterations_count: Number of optimizer iterations
2. Apache Kafka Integration¶
Schema Registry Automatic Versioning¶
The Schema Registry provides several key capabilities:
- Automatic Version Management: Each schema change creates a new version
- Compatibility Mode: Currently set to
NONE(unrestricted changes in development) - Schema Evolution: Supports adding/removing fields with default values
- Centralized Schema Store: Single source of truth for data structures
Schema Compatibility
The system uses NONE compatibility mode in the connector configuration, allowing unrestricted schema evolution. For production, consider BACKWARD or FULL compatibility. See the Confluent Schema Evolution docs for details.
Kafka Configuration Parameters¶
@dataclass
class ProducerConfig:
servers: str # Kafka bootstrap servers
topic: str # Target topic name
security: SecurityConfig
retries: int = 3 # Application-level retries
retry_delay: int = 2 # Delay between retries (seconds)
kafka_retries: int = 5 # Kafka internal retries
acks: str = 'all' # Acknowledgment level
timeout: int = 10 # Producer timeout (seconds)
Kafka runs in KRaft mode.
kafka:
image: bitnami/kafka:latest
environment:
- KAFKA_CFG_NODE_ID=0
- KAFKA_CFG_PROCESS_ROLES=controller,broker
- KAFKA_CFG_CONTROLLER_QUORUM_VOTERS=0@kafka:9093
- KAFKA_CFG_LISTENERS=PLAINTEXT://kafka:9092,CONTROLLER://:9093,EXTERNAL://0.0.0.0:9094
- KAFKA_CFG_ADVERTISED_LISTENERS=PLAINTEXT://kafka:9092,EXTERNAL://localhost:9094
- KAFKA_CFG_LISTENER_SECURITY_PROTOCOL_MAP=CONTROLLER:PLAINTEXT,PLAINTEXT:PLAINTEXT,EXTERNAL:PLAINTEXT
- KAFKA_CFG_CONTROLLER_LISTENER_NAMES=CONTROLLER
- KAFKA_CFG_INTER_BROKER_LISTENER_NAME=PLAINTEXT
Parallel Version Support¶
Allows multiple versions of the simulation module to run in parallel (e.g., when testing new functionality) without the risk of data structure conflicts. Kafka buffers results from multiple parallel simulator instances, ensuring transmission reliability in case of temporary consumer unavailability.
sequenceDiagram
participant P1 as Producer v1
participant P2 as Producer v2
participant K as Kafka
participant SR as Schema Registry
participant C as Consumer
P1->>SR: Register Schema v1
SR-->>P1: Schema ID: 101
P2->>SR: Register Schema v2
SR-->>P2: Schema ID: 102
par Parallel Production
P1->>K: Publish to vqe_decorated_result_v1
P2->>K: Publish to vqe_decorated_result_v2
end
C->>K: Subscribe to vqe_decorated_result_.*
K->>C: Deliver messages from v1
K->>C: Deliver messages from v2
Note over C: Consumer handles both versionsUse Cases:
- A/B Testing: Run different algorithm configurations simultaneously
- Blue-Green Deployments: Deploy new version without stopping old one
- Canary Releases: Gradually roll out schema changes
- Development/Production Isolation: Separate topics for different environments
3. MinIO Integration¶
S3-Compatible Object Storage¶
For persistent data storage, the system uses MinIO - an object storage compatible with the Amazon S3 API. Data is automatically transferred from Kafka to MinIO using the Kafka Connect S3 Sink connector.
graph LR
K[Kafka Topics] -->|Stream| KC[Kafka Connect]
KC <-->|Validate| SR[Schema Registry]
KC -->|S3 PutObject| MINIO[MinIO<br/>local-vqe-results]
style K fill:#ffe082,color:#000
style KC fill:#ffe082,color:#000
style SR fill:#ffe082,color:#000
style MINIO fill:#b39ddb,color:#311b92Kafka Connect S3 Sink Connector Configuration¶
The following configuration defines how data is stored, the serialization format, and connection parameters to MinIO:
{
"name": "minio-sink",
"config": {
"connector.class": "io.confluent.connect.s3.S3SinkConnector",
"tasks.max": "1",
"topics.regex": "vqe_decorated_result_.*",
"refresh.topics.enabled": "true",
"topics.dir": "experiments",
"directory.delim": "/",
"s3.bucket.name": "local-vqe-results",
"store.url": "http://minio:9000",
"s3.region": "us-east-1",
"s3.path.style.access": "true",
"s3.access.key.id": "admin",
"s3.secret.access.key": "admin123",
"s3.part.size": "5242880",
"s3.retry.backoff.ms": "1000",
"flush.size": "1",
"schema.compatibility": "NONE",
"storage.class": "io.confluent.connect.s3.storage.S3Storage",
"format.class": "io.confluent.connect.s3.format.avro.AvroFormat",
"key.converter": "io.confluent.connect.avro.AvroConverter",
"value.converter": "io.confluent.connect.avro.AvroConverter",
"key.converter.schema.registry.url": "http://schema-registry:8081",
"value.converter.schema.registry.url": "http://schema-registry:8081",
"errors.tolerance": "all",
"errors.log.enable": "true",
"errors.log.include.messages": "true"
}
}
Connector Configuration
Key project-specific choices: flush.size: 1 for immediate write-through, schema.compatibility: NONE for unrestricted schema evolution, and s3.path.style.access: true required for MinIO. See the Confluent S3 Sink Connector docs for parameter details.
Directory Structure in MinIO¶
The connector creates the following directory structure:
s3://local-vqe-results/
└── experiments/
├── vqe_decorated_result_v1/
│ ├── partition=0/
│ │ ├── vqe_decorated_result_v1+0+0000000000.avro
│ │ ├── vqe_decorated_result_v1+0+0000000001.avro
│ │ └── vqe_decorated_result_v1+0+0000000002.avro
│ ├── partition=1/
│ │ └── vqe_decorated_result_v1+1+0000000000.avro
│ └── partition=2/
│ └── vqe_decorated_result_v1+2+0000000000.avro
└── vqe_decorated_result_v2/
├── partition=0/
│ └── vqe_decorated_result_v2+0+0000000000.avro
└── partition=1/
└── vqe_decorated_result_v2+1+0000000000.avro
File Naming Convention:
Example: vqe_decorated_result_v1+0+0000000000.avro
- Topic:
vqe_decorated_result_v1 - Partition:
0 - Start Offset:
0000000000
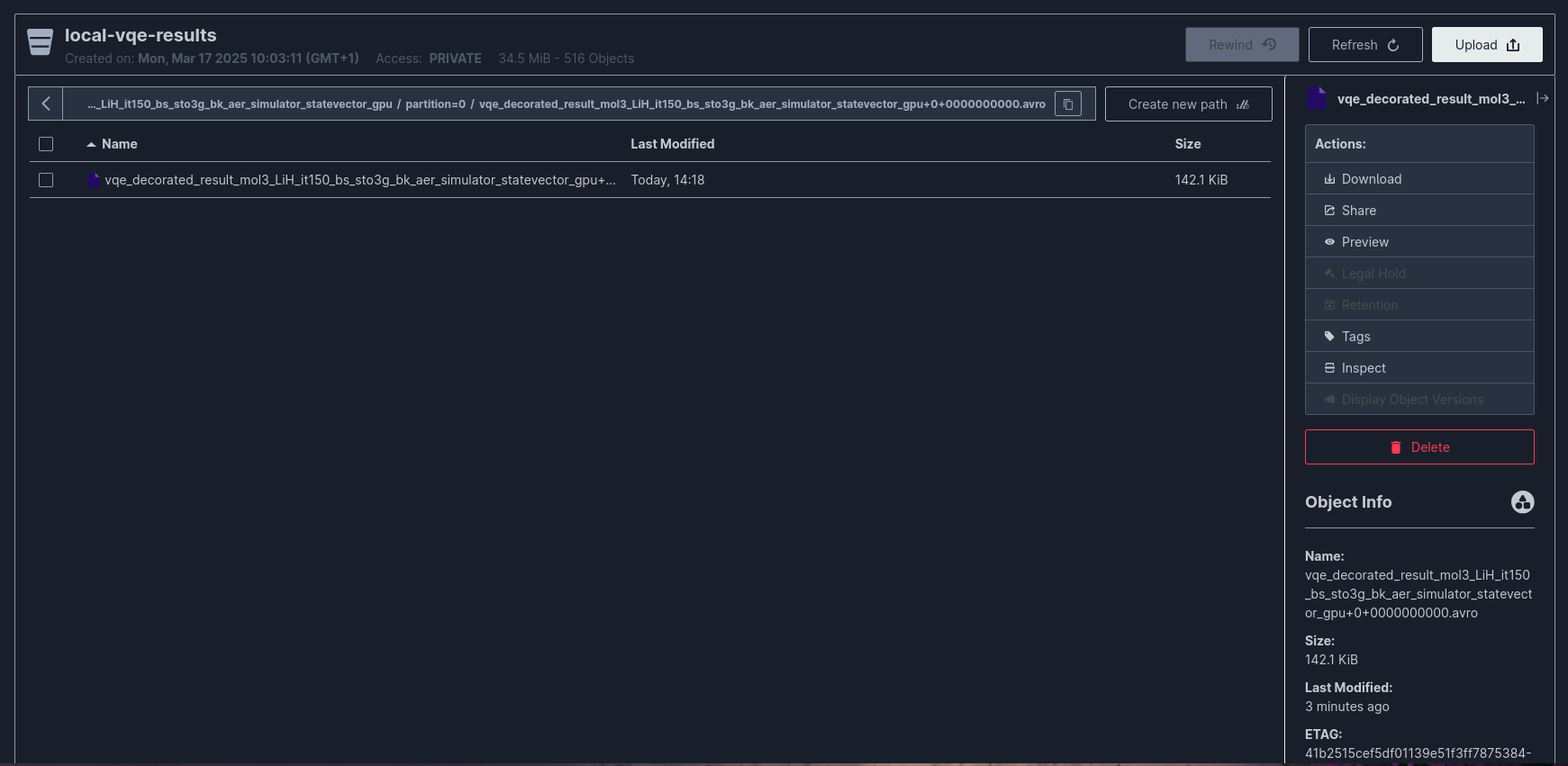
Data Consistency Guarantees¶
The connector configuration ensures data consistency through:
| Feature | Configuration | Guarantee |
|---|---|---|
| Exactly-Once Semantics | enable.idempotence: true (producer) |
No duplicate writes |
| Durability | acks: all (producer) |
Data persisted to all replicas |
| Error Handling | errors.tolerance: all |
Failed messages logged but processing continues |
| Schema Validation | Schema Registry integration | Only valid messages written to MinIO |
4. Apache Airflow Orchestration¶
Workflow Orchestration¶
Apache Airflow orchestrates the data processing workflow by managing the processing schedule. In this project, Airflow runs a daily incremental processing task (schedule_interval=timedelta(days=1)) that delegates computations to Apache Spark.
graph LR
subgraph "Airflow"
DAG["Feature Processing<br/>DAG"]
VARS["Variables<br/>MinIO credentials · Spark settings"]
DAG -->|reads| VARS
end
SPARK[Spark Cluster]
subgraph "Storage"
MINIO[(MinIO<br/>Raw Iceberg tables)]
ICEBERG[(MinIO<br/>Feature tables)]
end
DAG -->|SparkSubmitOperator| SPARK
SPARK -->|read| MINIO
SPARK -->|write| ICEBERG
style DAG fill:#90caf9,color:#0d47a1
style SPARK fill:#a5d6a7,color:#1b5e20
style MINIO fill:#ffe082,color:#e65100
style ICEBERG fill:#ffe082,color:#e65100DAG Definition¶
The DAG (Directed Acyclic Graph) defines the task for processing quantum experiment results into ML features:
with DAG(
'quantum_feature_processing',
default_args={**default_args, 'retries': 3, 'retry_delay': timedelta(minutes=20)},
description='Process quantum experiment data into ML feature tables',
schedule_interval=timedelta(days=1),
start_date=datetime(2025, 1, 1),
catchup=False,
tags=['quantum', 'ML', 'features', 'processing', 'Apache Spark'],
on_success_callback=send_success_email,
) as dag:
quantum_simulation_results_processing = SparkSubmitOperator(
task_id='run_quantum_processing',
application='/opt/airflow/dags/scripts/quantum_incremental_processing.py',
conn_id='spark_default',
name='quantum_feature_processing',
conf={
'spark.master': Variable.get('SPARK_MASTER'),
'spark.app.name': Variable.get('APP_NAME'),
'spark.hadoop.fs.s3a.endpoint': Variable.get('S3_ENDPOINT'),
'spark.hadoop.fs.s3a.access.key': Variable.get('MINIO_ACCESS_KEY'),
'spark.hadoop.fs.s3a.secret.key': Variable.get('MINIO_SECRET_KEY'),
'spark.hadoop.fs.s3a.path.style.access': 'true',
'spark.hadoop.fs.s3a.connection.ssl.enabled': 'false',
'spark.jars.packages': (
'org.slf4j:slf4j-api:2.0.17,'
'commons-codec:commons-codec:1.18.0,'
'com.google.j2objc:j2objc-annotations:3.0.0,'
'org.apache.spark:spark-avro_2.12:3.5.5,'
'org.apache.hadoop:hadoop-aws:3.3.1,'
'org.apache.hadoop:hadoop-common:3.3.1,'
'org.apache.iceberg:iceberg-spark-runtime-3.5_2.12:1.4.2'
),
},
env_vars={
'MINIO_ACCESS_KEY': Variable.get('MINIO_ACCESS_KEY'),
'MINIO_SECRET_KEY': Variable.get('MINIO_SECRET_KEY'),
'S3_ENDPOINT': Variable.get('S3_ENDPOINT'),
'S3_BUCKET': Variable.get('S3_BUCKET'),
'S3_WAREHOUSE': Variable.get('S3_WAREHOUSE'),
'SPARK_MASTER': Variable.get('SPARK_MASTER'),
},
verbose=True,
)
The DAG runs daily with catchup=False (no backfill), retries up to 3 times with 20-minute delays, and sends email notifications on success/failure. For Airflow scheduling and retry configuration details, see the Apache Airflow docs.
The SparkSubmitOperator delegates to the Spark cluster. All connection parameters (MinIO credentials, Spark master URL, S3 endpoints) are managed through Airflow Variables, initialized from the DEFAULT_CONFIG dict in quantum_incremental_processing.py and environment variables.
Default Configuration Values (from DEFAULT_CONFIG in the processing script):
DEFAULT_CONFIG = {
'S3_ENDPOINT': 'http://minio:9000',
'SPARK_MASTER': 'spark://spark-master:7077',
'S3_BUCKET': 's3a://local-vqe-results/experiments/',
'S3_WAREHOUSE': 's3a://local-features/warehouse/',
'APP_NAME': 'Quantum Pipeline Feature Processing',
}
DAG Execution Flow¶
sequenceDiagram
participant S as Airflow Scheduler
participant E as Airflow Executor
participant SSO as SparkSubmitOperator
participant SM as Spark Master
participant SW as Spark Workers
participant M as MinIO
participant I as Iceberg
Note over S: Cron: Daily at 00:00
S->>E: Schedule quantum_feature_processing
E->>SSO: Execute run_quantum_processing
SSO->>SSO: Read Airflow Variables
SSO->>SM: Submit PySpark job
SM->>SW: Distribute tasks
par Read Raw Data
SW->>M: Read Avro files from experiments/
end
SW->>SW: Transform to feature tables
par Write Features
SW->>I: Write to Iceberg tables
end
I->>I: Create snapshot
I->>M: Persist Parquet files
SW->>SM: Report completion
SM->>SSO: Job finished
SSO->>E: Task success
E->>S: Update DAG state
alt Success
S->>S: Send success email
else Failure
S->>S: Retry (up to 3 times)
S->>S: Send failure email
endThe Airflow web UI is accessible at http://airflow:8084 for monitoring DAG execution, task status, and logs.
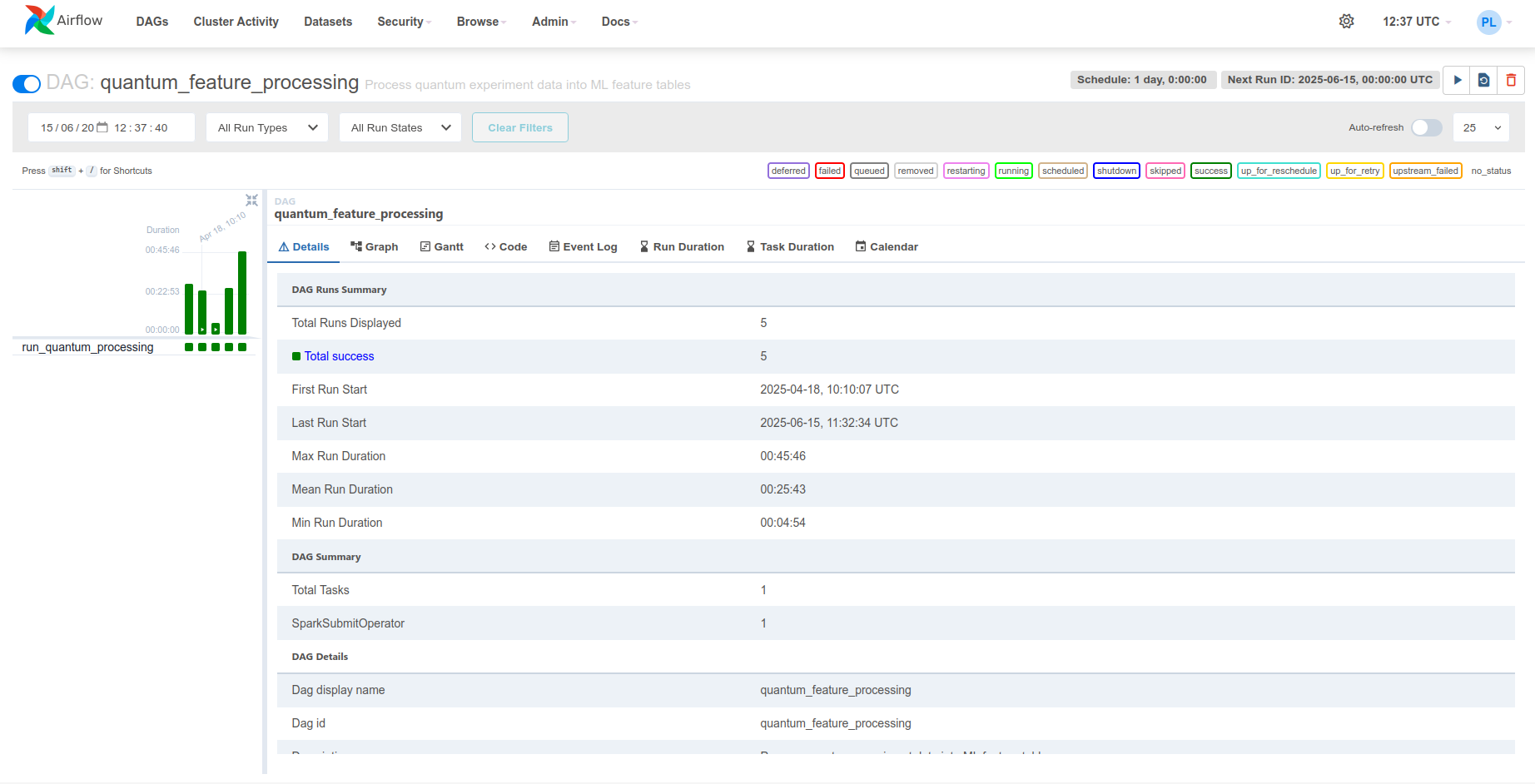
5. Incremental Processing¶
Overview¶
The system implements incremental processing, meaning that only new data since the last run is processed. This eliminates the need to repeatedly process the same results and significantly reduces execution time and computational resource usage.
Apache Iceberg provides a mechanism for tracking the state of processed data through a metadata layer. Iceberg stores information about data snapshots, enabling Spark to efficiently detect which files in MinIO have already been processed and which are new.
graph LR
RAW[(MinIO<br/>Raw Avro files)]
META[Iceberg Metadata<br/>Last processed snapshot]
subgraph "Spark"
FILTER[Identify new files]
PROC[Compute features]
FILTER -->|new files only| PROC
end
FEAT[(MinIO<br/>Feature tables)]
RAW -->|list files| FILTER
META -->|last snapshot| FILTER
PROC --> FEAT
PROC -->|update snapshot| META
style FILTER fill:#ffe082,color:#000
style RAW fill:#90caf9,color:#0d47a1
style FEAT fill:#a5d6a7,color:#1b5e20
style META fill:#b39ddb,color:#311b926-Step Processing Workflow¶
The incremental processing workflow consists of the following steps:
Step 1: Create Spark Session with MinIO Configuration¶
def create_spark_session(config=None):
"""Create and configure a Spark session with Iceberg and S3 support."""
if config is None:
config = DEFAULT_CONFIG
validate_environment()
access_key = os.environ.get('MINIO_ACCESS_KEY')
secret_key = os.environ.get('MINIO_SECRET_KEY')
return (
SparkSession.builder
.appName(config.get('APP_NAME'))
.master(config.get('SPARK_MASTER'))
.config('spark.jars.packages', (
'org.apache.spark:spark-avro_2.12:3.5.5,'
'org.apache.hadoop:hadoop-aws:3.3.1,'
'org.apache.iceberg:iceberg-spark-runtime-3.5_2.12:1.4.2'
))
.config('spark.sql.extensions',
'org.apache.iceberg.spark.extensions.IcebergSparkSessionExtensions')
.config('spark.sql.catalog.quantum_catalog',
'org.apache.iceberg.spark.SparkCatalog')
.config('spark.sql.catalog.quantum_catalog.type', 'hadoop')
.config('spark.sql.catalog.quantum_catalog.warehouse',
config.get('S3_WAREHOUSE'))
.config('spark.hadoop.fs.s3a.impl',
'org.apache.hadoop.fs.s3a.S3AFileSystem')
.config('spark.hadoop.fs.s3a.access.key', access_key)
.config('spark.hadoop.fs.s3a.secret.key', secret_key)
.config('spark.hadoop.fs.s3a.endpoint', config.get('S3_ENDPOINT'))
.config('spark.hadoop.fs.s3a.path.style.access', 'true')
.config('spark.hadoop.fs.s3a.connection.ssl.enabled', 'false')
.config('spark.sql.adaptive.enabled', 'true')
.config('spark.sql.shuffle.partitions', '200')
.getOrCreate()
)
The session configures the quantum_catalog Iceberg catalog backed by a Hadoop warehouse on MinIO (s3a://local-features/warehouse/), with adaptive query execution enabled and 200 shuffle partitions.
Step 2: Initialize or Read Iceberg Metadata¶
def list_available_topics(spark, bucket_path):
"""List available topics from the storage."""
fs = spark._jvm.org.apache.hadoop.fs.FileSystem.get(
spark._jvm.java.net.URI.create(bucket_path),
spark._jsc.hadoopConfiguration()
)
path = spark._jvm.org.apache.hadoop.fs.Path(bucket_path)
if fs.exists(path) and fs.isDirectory(path):
return [f.getPath().getName() for f in fs.listStatus(path) if f.isDirectory()]
return []
Iceberg Metadata Structure:
s3a://local-features/warehouse/
└── quantum_features/
├── vqe_results/
│ ├── metadata/
│ │ ├── v1.metadata.json
│ │ ├── v2.metadata.json
│ │ ├── snap-1234567890.avro
│ │ └── snap-1234567891.avro
│ └── data/
│ ├── part-00000.parquet
│ └── part-00001.parquet
└── vqe_iterations/
├── metadata/
└── data/
Step 3: Filter Data - Select Only New Files¶
def identify_new_records(spark, new_data_df, table_name, key_columns):
"""
Identifies records in new_data_df that don't exist in the target table.
Uses DataFrame operations for better performance.
"""
# If table does not exist, every record is new
if not spark.catalog.tableExists(f'quantum_catalog.quantum_features.{table_name}'):
return new_data_df
# Retrieve existing keys
existing_keys = spark.sql(
f'SELECT DISTINCT {", ".join(key_columns)} '
f'FROM quantum_catalog.quantum_features.{table_name}'
)
# If table exists but has no records - return all records
if existing_keys.isEmpty():
return new_data_df
# Anti-join to find new records
new_with_marker = new_data_df.select(*key_columns).distinct().withColumn('is_new', lit(1))
existing_with_marker = existing_keys.withColumn('exists', lit(1))
joined = new_with_marker.join(existing_with_marker, on=key_columns, how='left')
new_keys = joined.filter(col('exists').isNull()).select(*key_columns)
# Join back to get full records
truly_new_data = new_data_df.join(new_keys, on=key_columns, how='inner')
return truly_new_data
Deduplication Logic:
graph LR
A[New Data<br/>experiment_id: 1,2,3,4] --> B{Table Exists?}
B -->|No| C[All Records are New<br/>1,2,3,4]
B -->|Yes| D[Existing Data<br/>experiment_id: 1,2]
D --> E[Left Anti-Join]
A --> E
E --> F[Truly New Records<br/>experiment_id: 3,4]
style C fill:#a5d6a7,color:#1b5e20
style F fill:#a5d6a7,color:#1b5e20Step 4: Process Data - Extract Features¶
def transform_quantum_data(df):
"""Transforms the original quantum data into various feature tables."""
base_df = add_metadata_columns(df, 'quantum_base_processing')
# Extract nested fields
base_df = base_df.select(
col('experiment_id'),
col('molecule_id'),
col('basis_set'),
col('vqe_result.initial_data').alias('initial_data'),
col('vqe_result.iteration_list').alias('iteration_list'),
col('vqe_result.minimum').alias('minimum_energy'),
# ... more fields
)
# Create specialized feature tables
return {
'molecules': extract_molecule_features(base_df),
'vqe_results': extract_vqe_features(base_df),
'vqe_iterations': extract_iteration_features(base_df),
'hamiltonian_terms': extract_hamiltonian_features(base_df),
# ... 9 total feature tables
}
Feature Table Transformations:
df_molecule = base_df.select(
col('experiment_id'),
col('molecule_id'),
col('molecule_data.symbols').alias('atom_symbols'),
col('molecule_data.coords').alias('coordinates'),
col('molecule_data.multiplicity').alias('multiplicity'),
col('molecule_data.charge').alias('charge'),
col('processing_timestamp'),
col('processing_date'),
)
df_vqe = base_df.select(
col('experiment_id'),
col('molecule_id'),
col('basis_set'),
col('initial_data.backend').alias('backend'),
col('initial_data.num_qubits').alias('num_qubits'),
col('initial_data.optimizer').alias('optimizer'),
col('minimum_energy'),
size(col('iteration_list')).alias('total_iterations'),
col('processing_date'),
)
Step 5: Write Processed Data in Parquet Format¶
def process_incremental_data(spark, new_data_df, table_name, key_columns,
partition_columns=None, comment=None):
"""Process the data incrementally - only data that wasn't processed yet."""
# Check if table exists
table_exists = spark.catalog._jcatalog.tableExists(
f'quantum_catalog.quantum_features.{table_name}'
)
if not table_exists:
# Create new table
writer = new_data_df.write.format('iceberg').option('write-format', 'parquet')
if partition_columns:
writer = writer.partitionBy(*partition_columns)
writer.mode('overwrite').saveAsTable(
f'quantum_catalog.quantum_features.{table_name}'
)
else:
# Append only new records
truly_new_data = identify_new_records(spark, new_data_df, table_name, key_columns)
if truly_new_data.count() == 0:
return None, 0
truly_new_data.write.format('iceberg') \
.option('write-format', 'parquet') \
.mode('append') \
.saveAsTable(f'quantum_catalog.quantum_features.{table_name}')
return version_tag, record_count
Data is written in Parquet columnar format via Iceberg.
Step 6: Update Iceberg Metadata - Create New Snapshot¶
# Create a tag for this version of the table
snapshot_id = spark.sql(
f'SELECT snapshot_id FROM quantum_catalog.quantum_features.{table_name}.snapshots '
f'ORDER BY committed_at DESC LIMIT 1'
).collect()[0][0]
processing_batch_id = new_data_df.limit(1).collect()[0]['processing_batch_id'].replace('-', '')
version_tag = f'v_{processing_batch_id}'
spark.sql(f"""
ALTER TABLE quantum_catalog.quantum_features.{table_name}
CREATE TAG {version_tag} AS OF VERSION {snapshot_id}
""")
Iceberg Snapshot Management:
graph TB
subgraph "Timeline"
S1[Snapshot 1<br/>2025-01-10 00:00<br/>Tag: v_initial]
S2[Snapshot 2<br/>2025-01-11 00:00<br/>Tag: v_incr_abc123]
S3[Snapshot 3<br/>2025-01-12 00:00<br/>Tag: v_incr_def456]
S4[Snapshot 4<br/>Current]
end
S1 -->|+100 records| S2
S2 -->|+50 records| S3
S3 -->|+75 records| S4
subgraph "Time Travel Queries"
Q1[SELECT * FROM table<br/>VERSION AS OF v_initial]
Q2[SELECT * FROM table<br/>TIMESTAMP AS OF '2025-01-11']
Q3[SELECT * FROM table<br/>FOR SYSTEM_TIME AS OF '2025-01-12']
end
Q1 -.->|Returns| S1
Q2 -.->|Returns| S2
Q3 -.->|Returns| S3
style S4 fill:#b39ddb,color:#311b92Time-Travel Queries:
-- Query data as of specific snapshot
SELECT * FROM quantum_catalog.quantum_features.vqe_results
VERSION AS OF 'v_incr_abc123';
-- Query data at specific timestamp
SELECT * FROM quantum_catalog.quantum_features.vqe_results
TIMESTAMP AS OF '2025-01-11 12:00:00';
-- Compare snapshots
SELECT current.*, previous.*
FROM quantum_catalog.quantum_features.vqe_results FOR SYSTEM_VERSION AS OF 12345 AS current
JOIN quantum_catalog.quantum_features.vqe_results FOR SYSTEM_VERSION AS OF 12344 AS previous
ON current.experiment_id = previous.experiment_id
WHERE current.minimum_energy != previous.minimum_energy;
Feature Tables Schema¶
The system creates 9 specialized feature tables:
Purpose: Molecular structure information
| Column | Type | Description |
|---|---|---|
| experiment_id | string | Unique experiment identifier |
| molecule_id | int | Molecule identifier |
| atom_symbols | array<string> | Chemical symbols |
| coordinates | array<array<double>> | 3D coordinates |
| multiplicity | int | Spin multiplicity |
| charge | int | Molecular charge |
| coordinate_units | string | Units (angstrom/bohr) |
| atomic_masses | array<double> | Atomic masses |
Partitioning: processing_date
Purpose: Quantum circuit ansatz configurations
| Column | Type | Description |
|---|---|---|
| experiment_id | string | Unique experiment identifier |
| molecule_id | int | Molecule identifier |
| basis_set | string | Basis set (e.g., sto3g) |
| ansatz | string | QASM3 circuit representation |
| ansatz_reps | int | Number of repetitions |
Partitioning: processing_date, basis_set
Purpose: Execution timing and performance
| Column | Type | Description |
|---|---|---|
| experiment_id | string | Unique experiment identifier |
| molecule_id | int | Molecule identifier |
| basis_set | string | Basis set |
| hamiltonian_time | double | Hamiltonian build time (s) |
| mapping_time | double | Mapping time (s) |
| vqe_time | double | VQE execution time (s) |
| total_time | double | Total time (s) |
| minimization_time | double | Optimizer time (s) |
Partitioning: processing_date, basis_set
Purpose: VQE optimization results
| Column | Type | Description |
|---|---|---|
| experiment_id | string | Unique experiment identifier |
| molecule_id | int | Molecule identifier |
| basis_set | string | Basis set |
| backend | string | Qiskit backend |
| num_qubits | int | Number of qubits |
| optimizer | string | Optimizer algorithm |
| minimum_energy | double | Ground state energy (Ha) |
| total_iterations | int | Number of iterations |
Partitioning: processing_date, basis_set, backend
Purpose: Initial variational parameters
| Column | Type | Description |
|---|---|---|
| parameter_id | string | Unique parameter identifier |
| experiment_id | string | Experiment identifier |
| molecule_id | int | Molecule identifier |
| parameter_index | int | Parameter position |
| initial_parameter_value | double | Initial value |
Partitioning: processing_date, basis_set
Purpose: Optimized variational parameters
| Column | Type | Description |
|---|---|---|
| parameter_id | string | Unique parameter identifier |
| experiment_id | string | Experiment identifier |
| molecule_id | int | Molecule identifier |
| parameter_index | int | Parameter position |
| optimal_parameter_value | double | Optimized value |
Partitioning: processing_date, basis_set
Purpose: Per-iteration optimization data
| Column | Type | Description |
|---|---|---|
| iteration_id | string | Unique iteration identifier |
| experiment_id | string | Experiment identifier |
| molecule_id | int | Molecule identifier |
| iteration_step | int | Iteration number |
| iteration_energy | double | Energy at this step (Ha) |
| energy_std_dev | double | Standard deviation |
Partitioning: processing_date, basis_set, backend
Purpose: Parameters at each iteration
| Column | Type | Description |
|---|---|---|
| parameter_id | string | Unique parameter identifier |
| iteration_id | string | Iteration identifier |
| experiment_id | string | Experiment identifier |
| iteration_step | int | Iteration number |
| parameter_value | double | Parameter value |
Partitioning: processing_date, basis_set
Purpose: Hamiltonian Pauli operator terms
| Column | Type | Description |
|---|---|---|
| term_id | string | Unique term identifier |
| experiment_id | string | Experiment identifier |
| molecule_id | int | Molecule identifier |
| term_label | string | Pauli string (e.g., "XYZI") |
| coeff_real | double | Real coefficient |
| coeff_imag | double | Imaginary coefficient |
Partitioning: processing_date, basis_set, backend
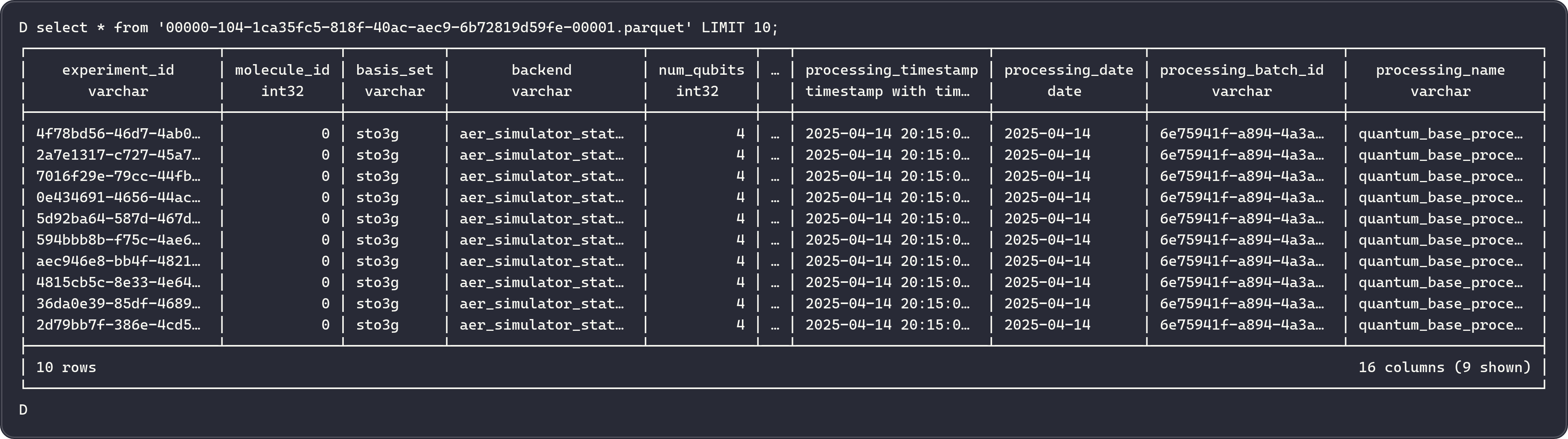
Efficient Filtering of Processed Files¶
The incremental processing system uses several optimization strategies:
1. Partition Pruning¶
# Only scan partitions for today
df = spark.read.format('iceberg') \
.load('quantum_catalog.quantum_features.vqe_results') \
.filter(col('processing_date') == current_date())
2. Predicate Pushdown¶
-- Filter pushed to storage layer
SELECT * FROM quantum_catalog.quantum_features.vqe_results
WHERE basis_set = 'sto3g' AND minimum_energy < -1.0;
3. Metadata-Only Queries¶
-- No data file reads required
SELECT COUNT(*) FROM quantum_catalog.quantum_features.vqe_results;
SELECT MIN(minimum_energy), MAX(minimum_energy)
FROM quantum_catalog.quantum_features.vqe_results;
4. File-Level Deduplication¶
Iceberg tracks individual data files, avoiding re-reading already processed files:
# Iceberg automatically skips files already in the table
new_files = set(current_files) - set(processed_files_in_snapshot)
Metadata Tracking Table¶
The system maintains a metadata table to track all processing runs:
CREATE TABLE IF NOT EXISTS quantum_catalog.quantum_features.processing_metadata (
processing_batch_id STRING,
processing_name STRING,
processing_timestamp TIMESTAMP,
processing_date DATE,
table_names ARRAY<STRING>,
table_versions ARRAY<STRING>,
record_counts ARRAY<BIGINT>,
source_data_info STRING
) USING iceberg
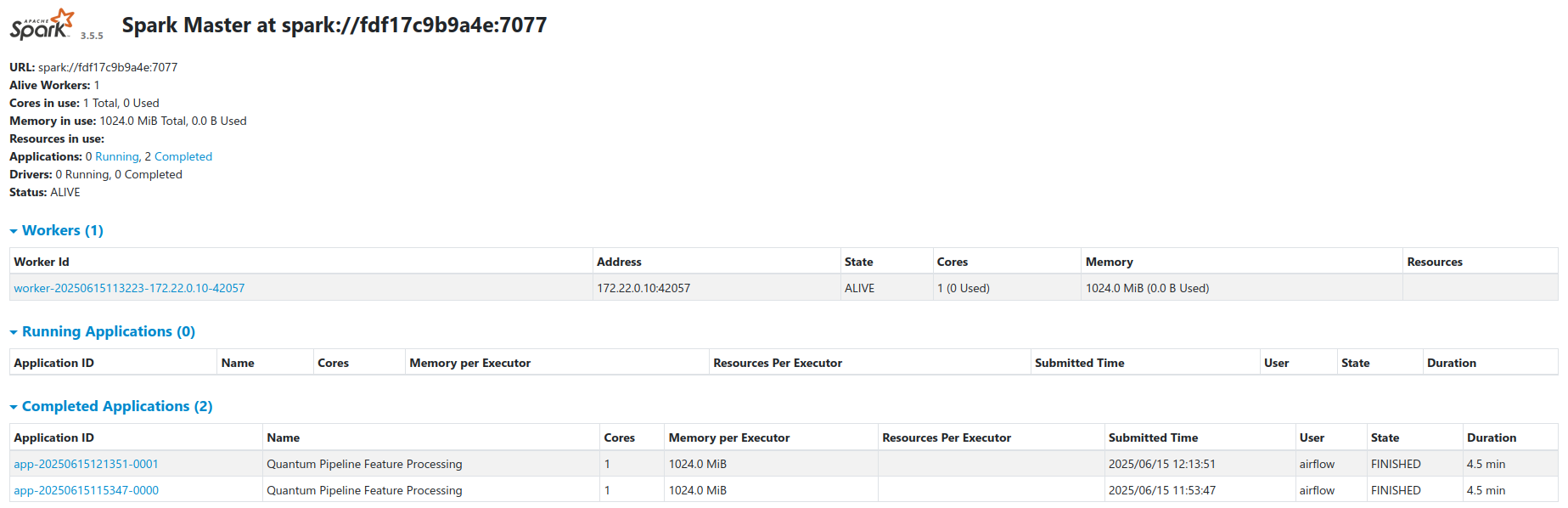
Deployment Architecture¶
Docker Compose Services¶
The complete platform is deployed using Docker Compose with the following services:
services:
# Data Streaming Layer - KRaft mode
kafka:
image: bitnami/kafka:latest
schema-registry:
image: confluentinc/cp-schema-registry:7.5.0
depends_on:
- kafka
kafka-connect:
image: confluentinc/cp-kafka-connect:7.5.0
depends_on:
- kafka
- schema-registry
- minio
# Storage Layer
minio:
image: minio/minio:latest
# Processing Layer
spark-master:
image: straightchlorine/quantum-pipeline:spark
spark-worker:
image: straightchlorine/quantum-pipeline:spark
depends_on:
- spark-master
# Orchestration Layer
postgres:
image: postgres:14
airflow-webserver:
image: straightchlorine/quantum-pipeline:airflow
depends_on:
- postgres
airflow-scheduler:
image: straightchlorine/quantum-pipeline:airflow
depends_on:
- postgres
# Quantum Simulation
quantum-pipeline:
image: straightchlorine/quantum-pipeline:latest
depends_on:
- kafka
- schema-registry
Monitoring and Observability¶
The system exports metrics to Prometheus via PushGateway (VQE execution metrics, resource utilization).
Grafana dashboards are accessible at http://grafana:3000 for VQE execution monitoring, data pipeline health.
For details on the monitoring stack configuration, see the Monitoring section.