GPU Acceleration¶
Quantum Pipeline leverages NVIDIA GPUs through Qiskit Aer's CUDA backend to accelerate quantum circuit simulation. This page covers the setup, configuration, and performance characteristics of GPU-accelerated execution.
Prerequisites¶
GPU acceleration requires an NVIDIA GPU (compute capability 6.0+), NVIDIA drivers (520+), CUDA Toolkit 11.8, NVIDIA Container Toolkit (1.14+), and Docker Engine (24.0+). For detailed installation steps, see the NVIDIA Container Toolkit installation guide.
cuQuantum Support
NVIDIA cuQuantum (cuStateVec, cuTensorNet) provides additional acceleration for quantum simulations but requires Ampere architecture or newer (compute capability 8.0+). The thesis experiments did not use cuQuantum due to hardware constraints.
GPU Configuration¶
Quantum Pipeline's GPU behavior is controlled through the backend configuration in
defaults.py:
'backend': {
'gpu': False, # Enable GPU acceleration
'gpu_opts': {
'device': 'GPU', # Target device ('GPU' or 'CPU')
'cuStateVec_enable': False, # Enable NVIDIA cuStateVec
'blocking_enable': False, # Reduce synchronization overhead
'batched_shots_gpu': True, # Enable shot parallelization for better GPU utilization
'shot_branching_enable': True, # Enable circuit branching optimization
'max_memory_mb': 5500, # GTX 1060: 6GB - 500MB buffer
},
}
These options are passed to Qiskit Aer's AerSimulator when GPU mode is enabled:
backend = AerSimulator(
method=self.backend_config.simulation_method,
**self.backend_config.gpu_opts,
noise_model=noise_model if noise_model else None,
)
Command-Line GPU Activation¶
Enable GPU acceleration from the command line with the --gpu flag:
python quantum_pipeline.py \
--file ./data/molecules.json \
--gpu \
--simulation-method statevector \
--max-iterations 150
Docker GPU Setup¶
Single Container¶
Run a GPU container with --gpus all:
docker run --rm --gpus all \
straightchlorine/quantum-pipeline:latest-gpu \
--file ./data/molecules.json \
--gpu \
--simulation-method statevector
To restrict to a specific GPU:
docker run --rm --gpus '"device=0"' \
straightchlorine/quantum-pipeline:latest-gpu \
--file ./data/molecules.json \
--gpu
Docker Compose¶
In Docker Compose, GPU access is configured through the deploy.resources.reservations
section:
deploy:
resources:
reservations:
devices:
- driver: nvidia
device_ids: ['0'] # Specific GPU by index
capabilities: [gpu]
Use count: all instead of device_ids to grant access to all available GPUs:
The thesis deployment assigns specific GPUs to specific containers using device_ids,
ensuring each pipeline instance has exclusive GPU access for accurate benchmarking.
Simulation Methods for GPU¶
Not all Qiskit Aer simulation methods support GPU acceleration:
| Method | GPU Support | Description |
|---|---|---|
statevector |
Yes | Full state vector simulation. Best for small-to-medium circuits. |
density_matrix |
Yes | Density matrix simulation. Supports noise models. |
tensor_network |
Yes (cuTensorNet) | Tensor network contraction. Requires cuQuantum. |
stabilizer |
No | Clifford circuit simulation. CPU only. |
unitary |
Yes | Full unitary matrix simulation. |
For the thesis experiments, statevector was used exclusively as it provides the
most direct benefit from GPU acceleration for VQE workloads. For the full comparison
table, GPU backend details, and memory scaling characteristics, see
Simulation Methods.
Performance Benchmarks¶
The thesis experiments evaluated GPU acceleration across six molecules of increasing complexity, comparing an Intel Core i5-8500 CPU against two NVIDIA GPUs.
Test Hardware¶
| Configuration | Processor / GPU | Memory |
|---|---|---|
| CPU Baseline | Intel Core i5-8500 (6 cores @ 3.00 GHz) | 16 GB RAM |
| GPU1 | NVIDIA GTX 1060 (1280 CUDA cores) | 6 GB VRAM |
| GPU2 | NVIDIA GTX 1050 Ti (768 CUDA cores) | 4 GB VRAM |
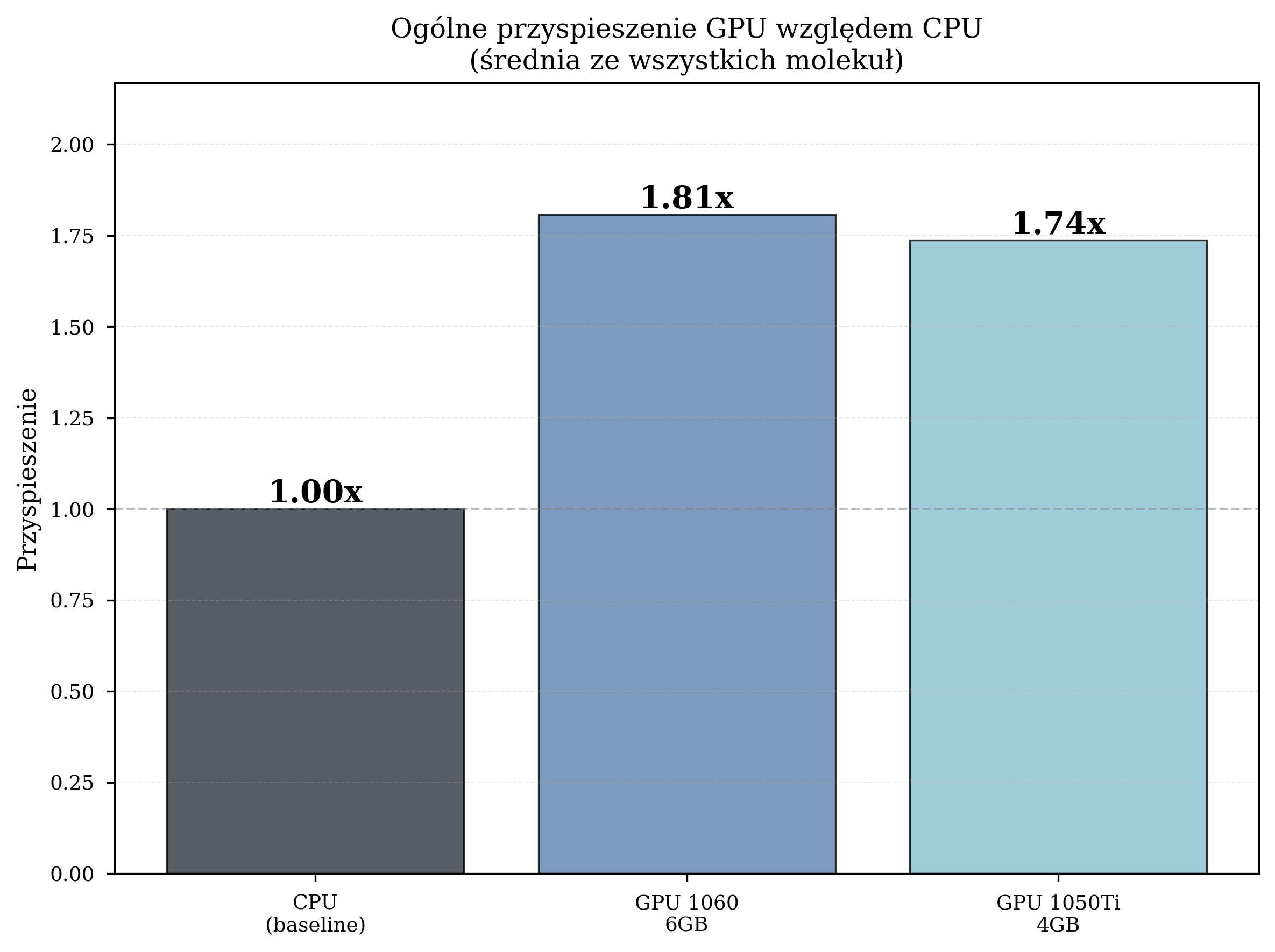
Overall Speedup (STO-3G, L-BFGS-B)¶
The primary thesis experiments used the STO-3G basis set with the L-BFGS-B gradient-based optimizer across six molecules of increasing complexity:
| Configuration | Avg. Time/Iteration | Total Iterations | Speedup |
|---|---|---|---|
| CPU (Intel i5-8500) | 4.259 s | 8,832 | 1.00x (baseline) |
| GPU GTX 1060 6GB | 2.357 s | 12,057 | 1.81x |
| GPU GTX 1050 Ti 4GB | 2.454 s | 10,871 | 1.74x |
GPU acceleration reduced the average iteration time by 42-45% compared to CPU execution. The GTX 1060, with more CUDA cores and VRAM, consistently outperformed the GTX 1050 Ti.
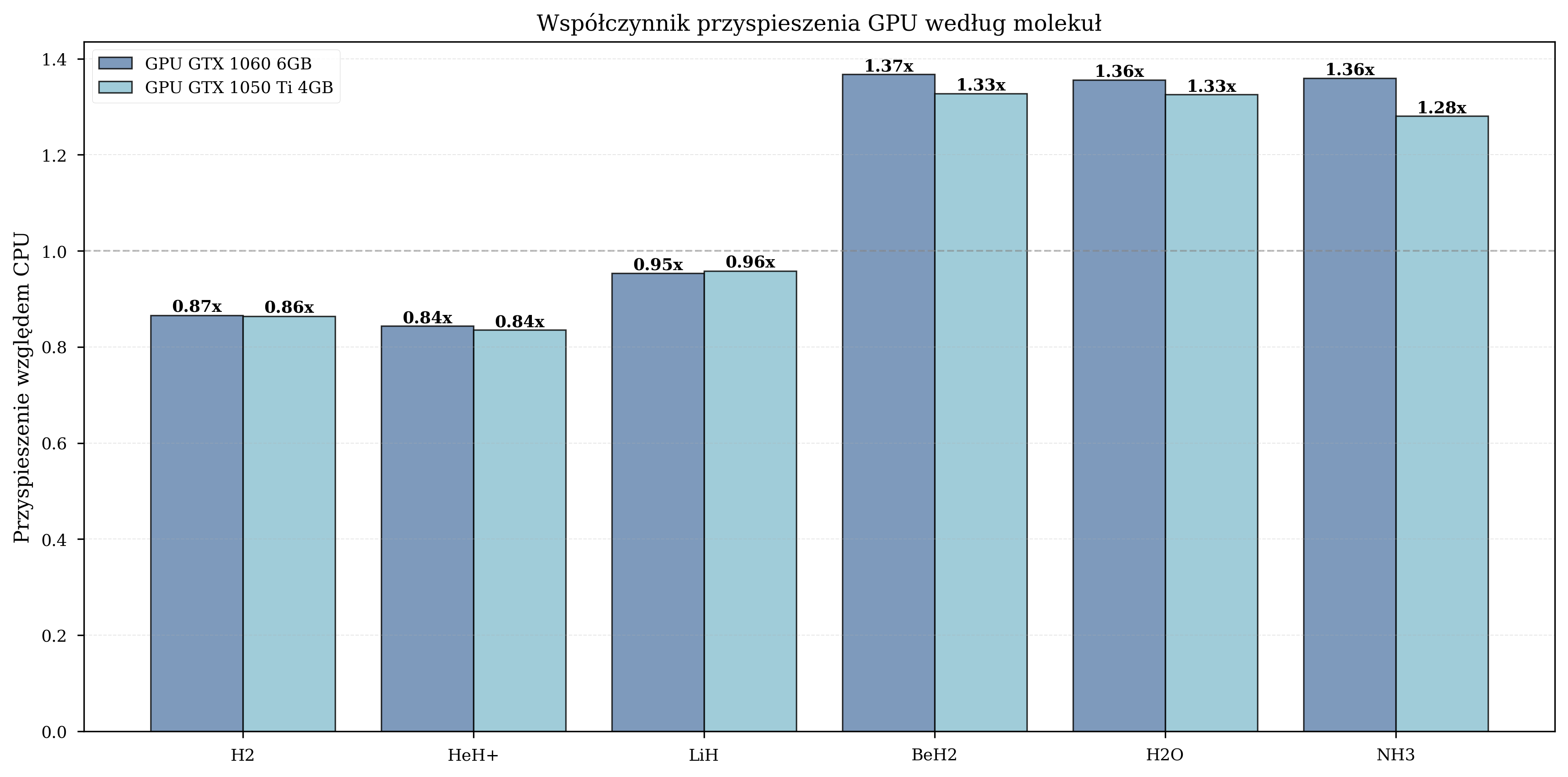
Speedup by Molecule¶
The benefit of GPU acceleration depends on the number of qubits in the simulation:
| Molecule | Qubits | GPU Speedup |
|---|---|---|
| H2 | 4 | 0.8-1.0x (GPU overhead dominates) |
| HeH+ | 4 | 0.9-1.1x (minimal benefit) |
| LiH | 8 | 1.5-1.6x (moderate) |
| BeH2 | 10 | 1.8-2.1x (peak benefit) |
| H2O | 12 | 1.3-1.4x (moderate) |
| NH3 | 12 | 1.3-1.4x (moderate) |
For small molecules (4 qubits), the overhead of CPU-GPU data transfer outweighs the computational benefit. Peak speedup occurs at 10 qubits (BeH2), where the state vector is large enough to saturate GPU parallelism. For larger molecules (12 qubits), the speedup remains significant but stabilizes as memory bandwidth becomes the limiting factor.
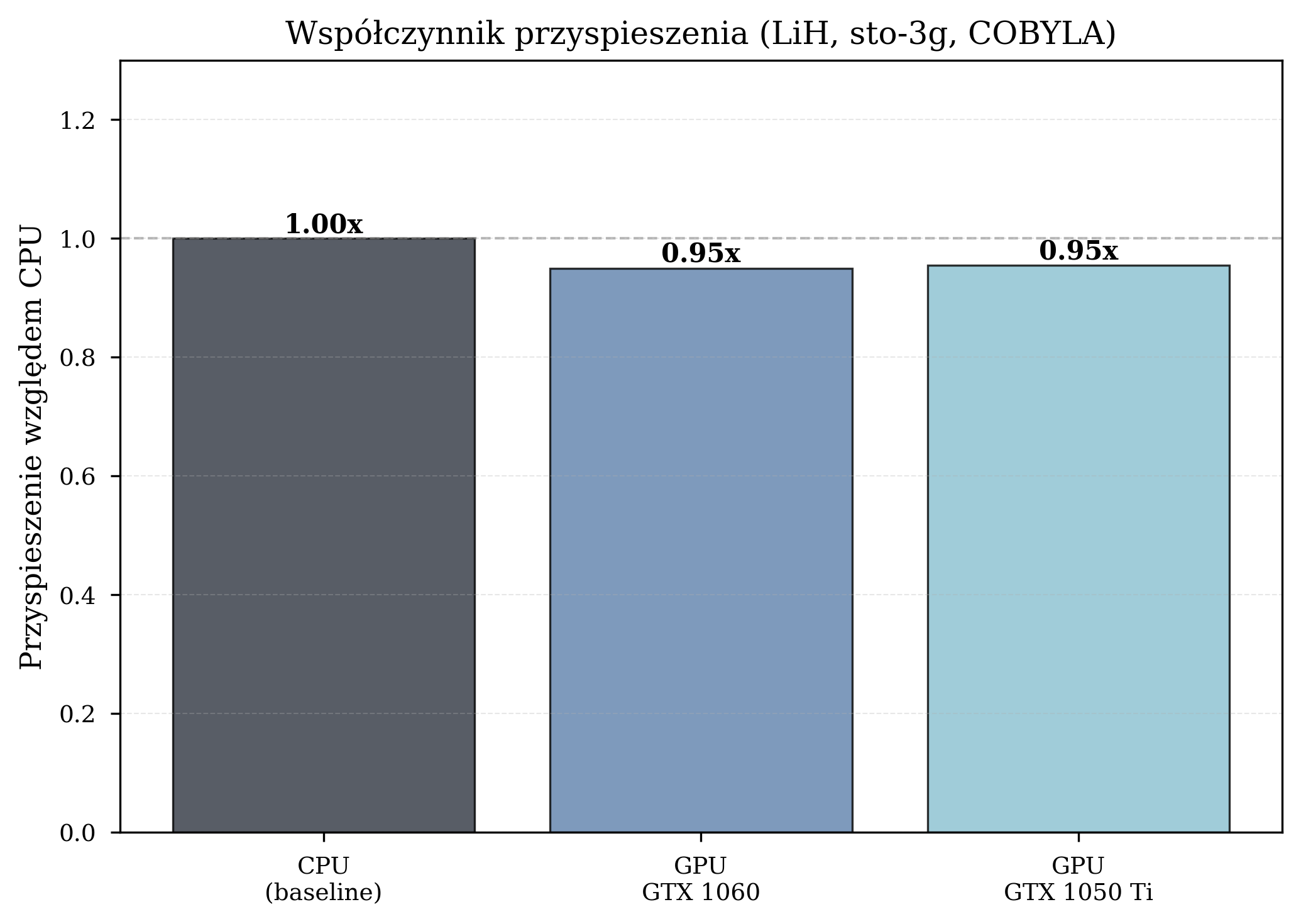
Impact of Basis Set Complexity¶
GPU acceleration benefits depend heavily on computational complexity. The figures below illustrate this: a small basis set with a derivative-free optimizer shows no GPU benefit, while a larger basis set with a gradient-based optimizer shows dramatic speedup.
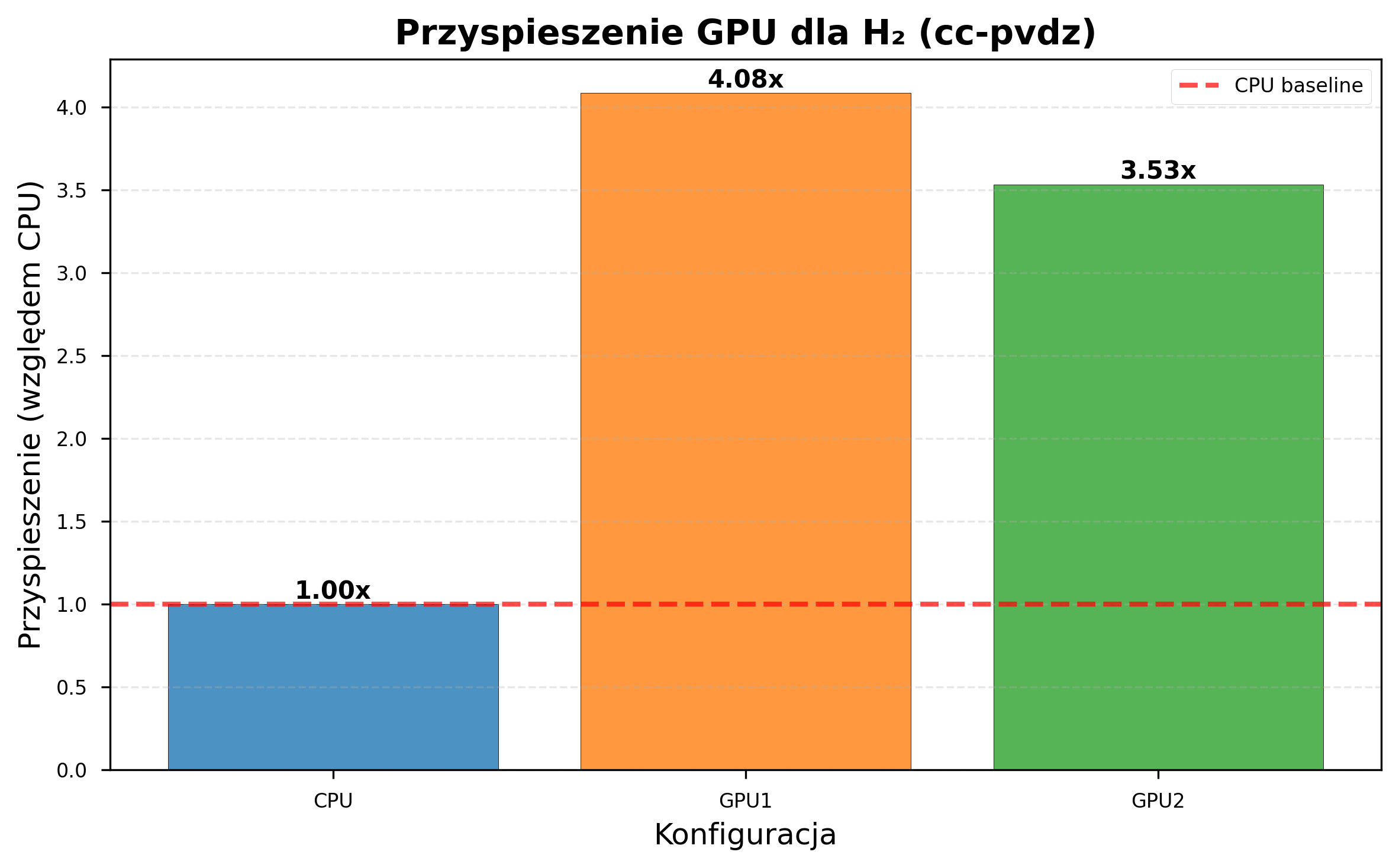
GPU acceleration becomes increasingly beneficial as problem complexity grows. With cc-pVDZ (206 s/iteration on CPU vs 51 s on GTX 1060), a simulation requiring 7 days on CPU completes in under 2 days on GPU.
Troubleshooting¶
For general GPU and Docker troubleshooting (driver installation, container toolkit configuration, runtime issues), see the NVIDIA Container Toolkit troubleshooting guide.
Out of Memory: The state vector size doubles with each additional qubit. For a
4 GB GPU, the practical limit is approximately 28 qubits with statevector
simulation.
Qiskit Aer GPU Build: Ensure the AER_CUDA_ARCH flag in the Dockerfile matches
your target GPU architecture. Mismatched flags produce builds that fail to compile or
fail at runtime with CUDA error: no kernel image is available.
References¶
- NVIDIA Container Toolkit Installation Guide
- NVIDIA cuQuantum Documentation
- Qiskit Aer GPU Simulation
- Bayraktar, H. et al. cuQuantum SDK: A High-Performance Library for Accelerating Quantum Science. arXiv:2308.01999 (2023)